This piece is a continuation of the article on the efficient and unique pooling methods used to reduce the number of trials required to test patients for COVID-19, called Tapestry. These innovative pooling methods were conceptualised and tested by Prof Manoj Gopalkrishnan and his team from IIT Bombay in association with NCBS (National Centre for Biological Sciences). These methods would cater to the requirement of quick and efficient testing posed by the huge population of India.

In the previous article, we had discussed a few algorithms used to pool and test samples. Initially, the team tried out an adaptive testing method similar to the binary search algorithm where they would split the samples and perform sequential testing. This involves the rejection of pools with negative results and then investigating in the pool with positive samples. But, this procedure proved to be a slow one as multiple tests are performed sequentially and it takes 3-4 hours to perform a single test. Next, they tried a non-adaptive testing method called Combinatorial Orthogonal Matching Pursuit (COMP). Since it’s a non-adaptive method it was possible to get results of all samples in one round of testing. The results were accurate when the ratio of positive samples in a set was less, but as soon as the ratio increased, this algorithm yielded many ‘false positives’.
To tackle the drawbacks of the methods discussed above, the team came up with ‘compressed sensing’ methods which yielded amazing results. In the compressed sensing methods, they could also determine ‘viral loads’ in a sample which indicates the severity of the infection. Now, let us delve into the technicalities of compressed sensing, its uniqueness and significance.
How does it work?
To go into detail about compressed sensing, we will have to start looking into the mathematics behind the pooling of samples. Here, we can begin by declaring certain variables used in the problem:
n – number of patients to be tested
m – number of tests performed
x – vector of n elements denoting the estimated viral load of each of the patients. A viral load of 0 indicates no infection. Also, since the number of COVID cases in the country is sparse across the population, we assume x to be a sparse vector (one with many 0s as the entries).
y – vector of m elements denoting the measured viral load of each pool of samples being tested. y can be thought of as a “given” variable to understand the problem better. The main thrust of the problem is to find the estimated viral load (‘x’) given the measured viral load (‘y’) using m tests.
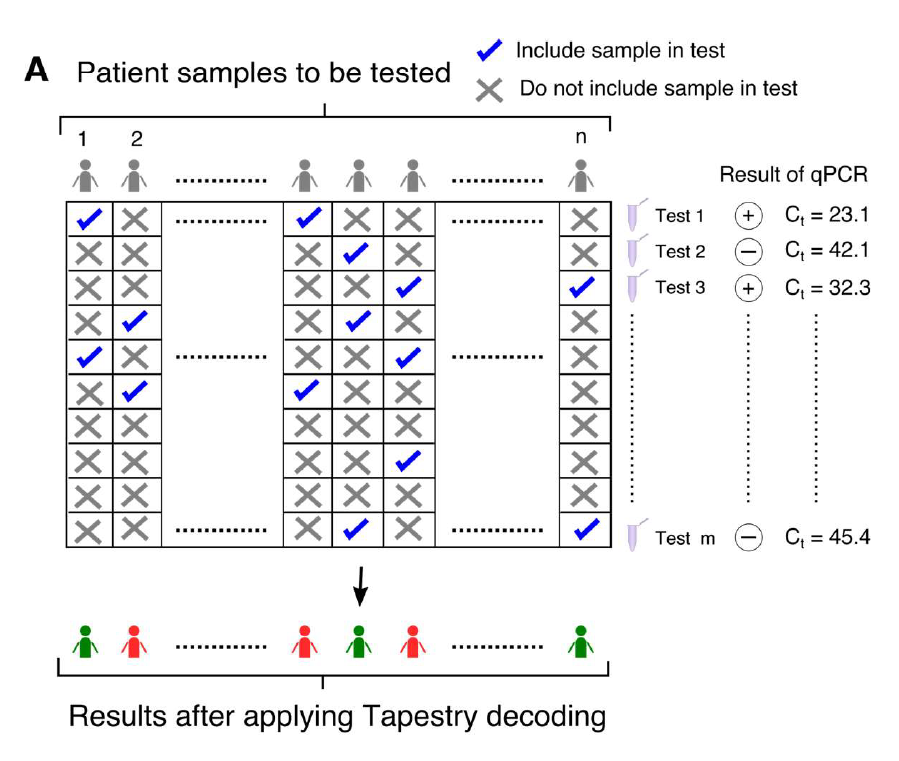
A – Binary mixing matrix of dimensions m*n. Aij = 1 implies that the jth sample is used in the ith test. Aij = 0 implies that the jth sample is not present in the ith test.
The equation to be solved is y = Ax.
|| y – Ax ||2: This function is used to determine how close the actual viral loads are to the calculated viral loads. The closer the error is to zero, the more accurate are our calculated viral loads.
Thus, we must have two objectives:
1) To find an x for which Ax is as close as possible to y.
2) Ensure that x is sparse, meaning that it has a large number of 0 entries.
Since the number of variables here exceeds the number of equations, we need to apply certain further constraints on the problem to estimate the viral loads.
There are a few methods that can be used to ensure the two objectives. Here, we can discuss the most successful ones so far.
Non Negative LASSO:
This method uses a cost function which penalises all the solutions which do not perform the above objectives. Thus the goal is to find a vector x which minimises the cost function, given by:
Jlasso = || y – Ax ||2 + || x ||
Here the mod (|| x ||) operator is the sum of the absolute values of x. This part of the cost function penalises all the values of x which have large entries. It also ensures the sparsity of the vector x.
The distance operator (|| y – Ax ||2) is the sum of squares of all the entries in the vector y – Ax. This part ensures that Ax is as close to y as possible. This ensures that the vector x is such that upon pooling the samples, we get results similar to those measured (in vector y).
NN LASSO was able to achieve a sensitivity of 99% and a specificity of 96% while being tested on computer-generated samples having one in ten positive cases (which is very similar to the ratio of positive samples in a set in India).
Non- Negative Orthogonal Matching:
NNOMP is a greedy approximation algorithm which tries to solve the optimisation problem given by || y – Ax ||2 < Ɛ, such that non-zero elements in x are minimum (this is done to impose the sparsity condition). The algorithm targets those samples which are most likely to contain the virus and assigns a viral load to it which would minimise || y – Ax ||2, a measure of the error of our calculated viral loads.
We keep assigning positive viral loads to elements in the vector x until the error passes below a threshold, Ɛ. In the next step, all the remaining entries in x are assigned 0, thus ensuring sparsity.
Although computationally expensive, this method has also been able to achieve surprising results. In computer-simulated samples, it showed a sensitivity of 95% and specificity of 98% in cases where one in 10 samples were positive.
How are they Different?
What makes the research done by the Tapestry team unique is the fact that having collaborated with the NCBS, they have been able to test their research on actual COVID samples. Rather than just produce algorithms to improve testing, they produced results in real-life lab situations, proving the value of their work beyond doubt.
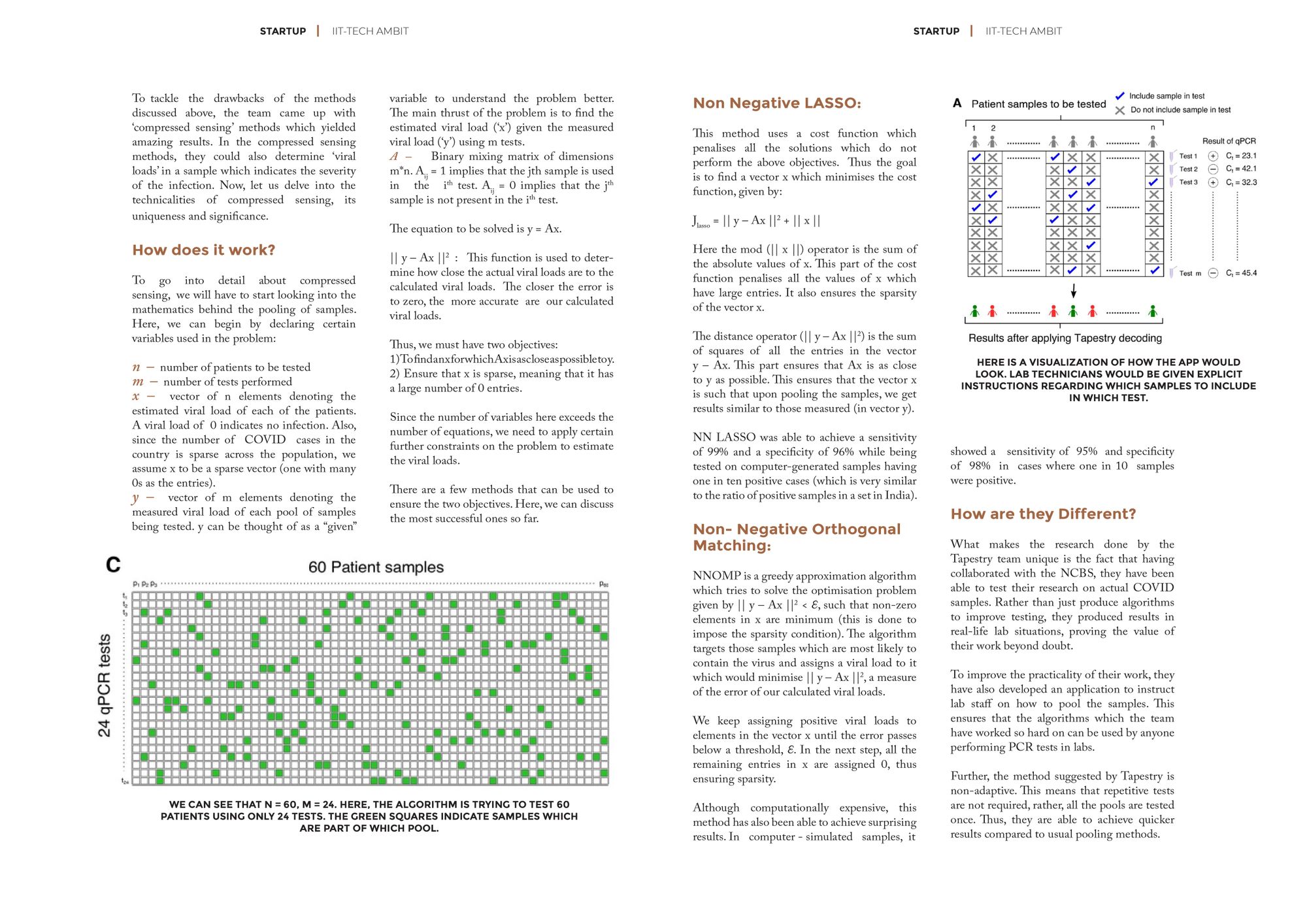
To improve the practicality of their work, they have also developed an application to instruct lab staff on how to pool the samples. This ensures that the algorithms which the team have worked so hard on can be used by anyone performing PCR tests in labs.
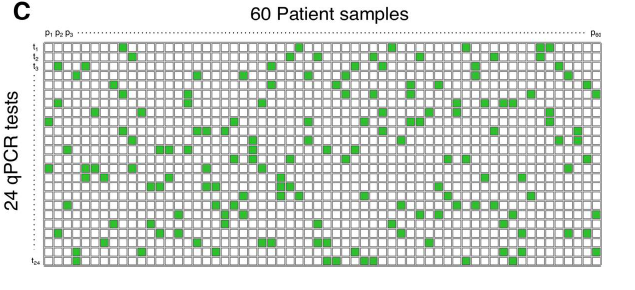
Further, the method suggested by Tapestry is non-adaptive. This means that repetitive tests are not required, rather, all the pools are tested once. Thus, they are able to achieve quicker results compared to usual pooling methods.
Keep an eye out for Team Tapestry
Currently, the team is working on various initiatives to test their pooling technique. From participating in international competitions such as Xprize, to securing government funding, team Tapestry is on its way to achieving fame on a global platform. They are also working on methods to make the entire testing process for COVID 19 completely automated. If successful, an automated COVID test would require no personnel, making it significantly safer and more efficient compared to the current testing methods.
In an overpopulated country with limited resources, implementation of compressed sensing methods would prove to be a boon. The testing rates would increase manifold, and it would be quite beneficial for us. Increased testing would mean more identification of positive cases. Thus, we can provide timely treatment to the affected people as well as isolate them to prevent the spread of the virus. To make this possible, the team is striving continuously to reach a near-perfect accuracy by trying out more such methods.
Team Tapestry is at the cusp of achieving greatness at several platforms, both nationally and internationally. Their success would ensure that limited testing rates would no longer serve as a hindrance to monitor the COVID spread in our country.




