Evolving technologies in all fields have demanded autonomous features over time. As visible to us, there are numbers to render right now- cars, self-navigating drones, military robots, underwater vehicles, and getting quite optimistic in enlisting all those names that ease up our technical chores and put a significant impact in increasing the productivity too.
Based on recent breakthroughs in the domains of deep learning and artificial intelligence, extensive and prompt regulatory developments are needed to specify the requirements from them and manage their deployment. And they all need unprecedented levels of safety and security, to overcome concerns about the potential negative impact of the new technology.
Such is the case in the identification of an indefectible drug or admixture of drugs while referring to a patient. Reasonably, we search for a reliable algorithm in providing us with the pieces of information on drugs that can counter the affiliated health issues.

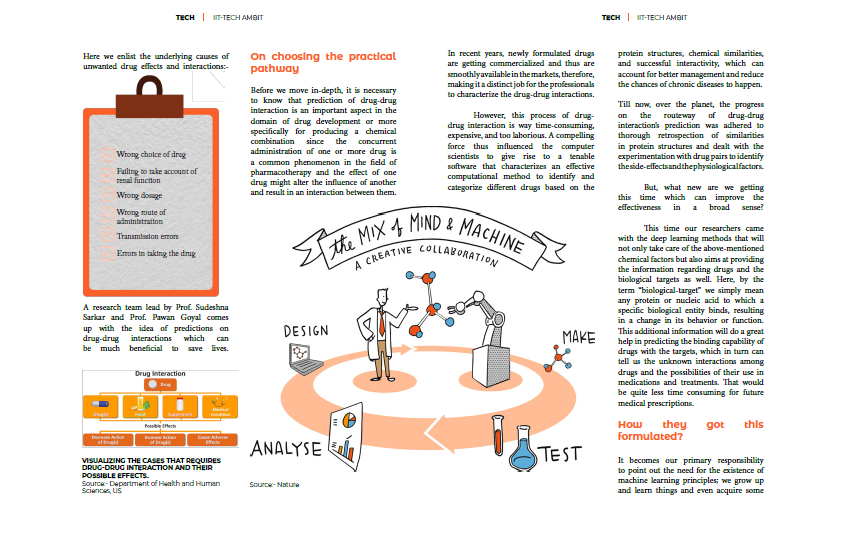
Interactions between drugs can lead to serious unwanted effects or a reduction in the therapeutic effects of some drug compositions. As we see, in elderly patients, there is a regular prescription of concurrent use of multiple medications, which the medical practitioners call Polypharmacy. It is not something to be necessarily considered as an ill-recommendation, about more than 40% of the older adults are relying on polypharmacy. But also it is a severe matter of concern since, in many instances, it can lead to adverse outcomes or reduced treatment effectiveness and sometimes even being more harmful than helpful or presenting too much risk for petty benefit. Here we enlist the underlying causes of unwanted drug effects and interactions:-
⁍Wrong choice of drug.
⁍Failing to take account of renal function
⁍Wrong dosage
⁍Wrong route of administration
⁍Transmission errors
⁍Errors in taking the drug
A research team lead by Prof. Sudeshna Sarkar and Prof. Pawan Goyal comes up with the idea of predictions on drug-drug interactions which can be much beneficial to save lives.
On choosing the practical pathway
Before we move in-depth, it is necessary to know that prediction of drug-drug interaction is an important aspect in the domain of drug development or more specifically for producing a chemical combination since the concurrent administration of one or more drug is a common phenomenon in the field of pharmacotherapy and the effect of one drug might alter the influence of another and result in an interaction between them. In recent years, newly formulated drugs are getting commercialized and thus are smoothly available in the markets, therefore, making it a distinct job for the professionals to characterize the drug-drug interactions.
However, this process of drug-drug interaction is way time-consuming, expensive, and too laborious. A compelling force thus influenced the computer scientists to give rise to a tenable software that characterizes an effective computational method to identify and categorize different drugs based on the protein structures, chemical similarities, and successful interactivity, which can account for better management and reduce the chances of chronic diseases to happen.

Till now, over the planet, the progress on the routeway of drug-drug interaction’s prediction was adhered to thorough retrospection of similarities in protein structures and dealt with the experimentation with drug pairs to identify the side-effects and the physiological factors.
But, what new are we getting this time which can improve the effectiveness in a broad sense?
This time our researchers came with the deep learning methods that will not only take care of the above-mentioned chemical factors but also aims at providing the information regarding drugs and the biological targets as well. Here, by the term “biological-target” we simply mean any protein or nucleic acid to which a specific biological entity binds, resulting in a change in its behavior or function. This additional information will do a great help in predicting the binding capability of drugs with the targets, which in turn can tell us the unknown interactions among drugs and the possibilities of their use in medications and treatments. That would be quite less time consuming for future medical prescriptions.
How did they get this formulated?
It becomes our primary responsibility to point out the need for the existence of machine learning principles; we grow up and learn things and even acquire some habits over time, but machines are not gifted with brains. They only follow those instructions that are put forward by the user, thus it became the need of the hour to train the machines priorly by passing some specific set of pieces of information which will vary according to the specified cases.

Machine learning is a subset of the larger field of artificial intelligence that provides systems the ability to automatically learn and improve from experience without being explicitly programmed. Machine learning focuses on the development of computer programs that can access data and use it to learn for themselves. The process of learning begins with observations or data, such as examples, direct experience, or instruction, to look for patterns in data and make better decisions in the future based on the examples that we provide. The primary aim is to allow the computers to learn automatically without human intervention or assistance and adjust actions accordingly.
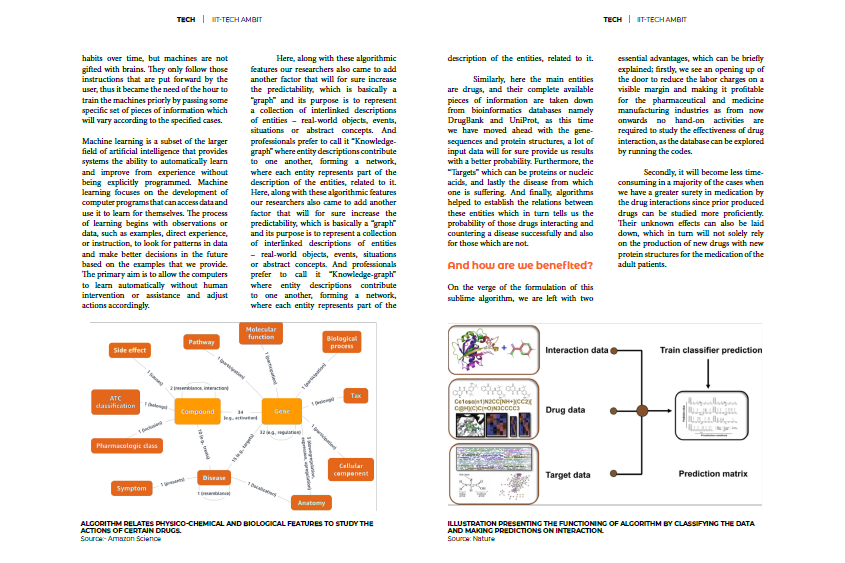
Here, along with these algorithmic features our researchers also came to add another factor that will for sure increase the predictability, which is basically a “graph” and its purpose is to represent a collection of interlinked descriptions of entities – real-world objects, events, situations or abstract concepts. And professionals prefer to call it “Knowledge-graph” where entity descriptions contribute to one another, forming a network, where each entity represents part of the description of the entities, related to it.
Similarly, here the main entities are drugs, and their complete available pieces of information are taken down from bioinformatics databases namely DrugBank and UniProt, as this time we have moved ahead with the gene-sequences and protein structures, a lot of input data will for sure provide us results with a better probability. Furthermore, the “Targets” which can be proteins or nucleic acids, and lastly the disease from which one is suffering. And finally, algorithms helped to establish the relations between these entities which in turn tells us the probability of those drugs interacting and countering a disease successfully and also for those which are not.
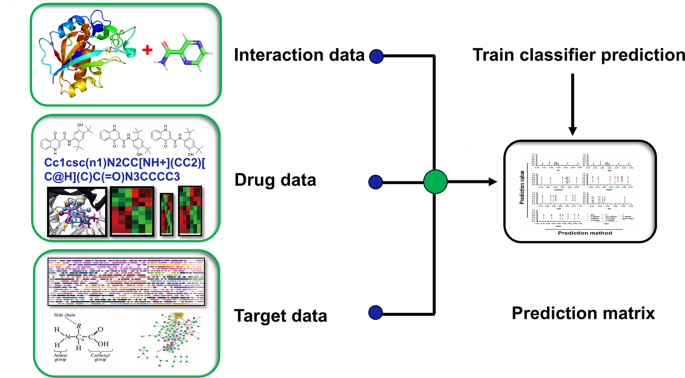
How are we benefited?
On the verge of the formulation of this sublime algorithm, we are left with two essential advantages, which can be briefly explained; firstly, we see an opening up of the door to reduce the labor charges on a visible margin and making it profitable for the pharmaceutical and medicine manufacturing industries as from now onwards no hand-on activities are required to study the effectiveness of drug interaction, as the database can be explored by running the codes.
Secondly, it will become less time-consuming in a majority of the cases when we have a greater surety in medication by the drug interactions since prior produced drugs can be studied more proficiently. Their unknown effects can also be laid down, which in turn will not solely rely on the production of new drugs with new protein structures for the medication of the adult patients.

